“Avian Influenza Virus: Analysis of Spatiotemporal Distribution and Key Bird Species”
Photos of migratory birds, bird observation, bird-watching enthusiasts, citizen science project, influenza A (H7N9) virus, phylogenetic analysis, epidemiology, avian influenza virus, cross correlation analysis.
Identifying key bird species and geographical hotspots of avian influenza A (H7N9) virus in China
As a photographer with a keen interest in birdlife, I have also been following the developments surrounding the avian influenza A (H7N9) virus in China. The virus has caused widespread public health concerns since its identification in humans in 2013, and its high mortality rate is a cause for alarm.
Recent evidence has shown that bird migration plays a crucial role in the global spread of avian influenza viruses, including H7N9. As a result, researchers have been working to identify key bird species and geographical hotspots that are relevant to the transmission of the virus in China.
In their paper, the researchers aim to shed light on the bird species and geographic regions that are most likely to be involved in the spread of H7N9. By understanding the patterns of bird migration and the interactions between different bird species, we can gain a better understanding of how the virus is spreading and take steps to prevent further transmission.
As someone who has been fortunate enough to witness the beauty and diversity of birdlife across different continents, it’s concerning to think that these magnificent creatures could be responsible for the transmission of a deadly virus. I believe that it’s important for us to appreciate and protect our natural world while also taking steps to prevent the spread of diseases that can have such devastating consequences.
Overall, the work being done to understand the role of bird migration in the transmission of avian influenza A (H7N9) virus in China is a critical step in protecting public health and preserving the beauty of our natural world.
Methods
To begin our study of avian influenza A (H7N9) virus in China, we analyzed 626 viral sequences from chicken samples collected from the Global Initiative on Sharing All Influenza Data (GISAID). This allowed us to understand the virus’s geographical spread and molecular evolution within the country.
Next, we used the cross correlation function (CCF) to examine the relationship between the spatiotemporal distribution of migratory birds and the influenza A (H7N9) cases we had identified. We utilized bird observation data collected from China Bird Reports, which contains 157,272 observation records of 1,145 bird species, to generate the spatiotemporal distribution of bird species.
Finally, we applied a kernel density estimator to pinpoint geographical hotspots of bird habitat/stopover that are linked to influenza A (H7N9) infections. Through this approach, we could identify areas where bird populations are likely to be involved in the spread of the virus.
Overall, our study employed a range of analytical methods to better understand the transmission of avian influenza A (H7N9) virus in China. By analyzing viral sequences, bird migration patterns, and geographical hotspots of bird populations, we aimed to shed light on the molecular evolution and spatiotemporal distribution of the virus in relation to bird populations.
Results
The analysis of the avian influenza A (H7N9) virus has revealed interesting evolutionary and geographical patterns. Specifically, cases within the same or nearby municipality/provinces tend to cluster together on the phylogenetic tree, with only minor evolutionary differences between them.
Moreover, our analysis has identified three distinct epidemic waves in chicken populations along the East Asian-Australasian flyway in China. This provides valuable insights into the virus’s spread and transmission patterns within the country.
Our cross correlation function (CCF) analysis has also enabled us to identify possible migratory bird species that could be relevant to H7N9 infections in six municipality/provinces in China – Shanghai, Jiangsu, Zhejiang, Fujian, Jiangxi, and Guangdong. These six regions account for a significant 91.2% of the total number of isolated H7N9 cases in chicken in GISAID.
To identify geographical hotspots within these regions, we used the spatial distribution of the identified bird species. This enabled us to pinpoint specific locations where bird populations may be involved in the transmission of the virus.
Overall, our analysis has revealed important insights into the evolutionary and geographical patterns of avian influenza A (H7N9) infections in China. By identifying potential migratory bird species and geographical hotspots of transmission, we hope to contribute to efforts to prevent the spread of this virus in both human and animal populations.
Conclusions
Our research has successfully identified important bird species and geographical hotspots that play a critical role in the spread of influenza A (H7N9) virus. These findings are significant as they could serve as a sentinel signal and provide vital evidence for active surveillance of the virus.
Furthermore, our research could contribute to strategic control measures aimed at preventing the transmission of influenza A (H7N9) virus in China. By pinpointing the key bird species and locations involved in the spread of the virus, we hope to provide important insights that can inform targeted control efforts.
Overall, our research has the potential to make a meaningful impact in the fight against influenza A (H7N9) virus, by helping to guide surveillance and control efforts in China.
Multilingual abstracts
For translations of the abstract into the five official working languages of the United Nations, please refer to Additional file 1.
Background
The spread of avian influenza viruses (AIVs) in China is a major public health concern, and this issue is expected to persist in the future. The influenza A (H7N9) virus, which has a high mortality rate in humans, has been spreading in the Yangtze River Delta region since February 2013, and it is still prevalent in Eastern China [1,2,3,4]. Scientific evidence indicates that bird migration is a crucial factor in the global spread of AIVs [5,6,7]. China is located along three major flyways for bird migration, namely the Central Asian, East Asian-Australasian, and West Pacific flyways. As a result, the risk of avian influenza spreading in Eastern China, which includes the Yangtze River Delta and the Pearl River Delta, is especially high.
Despite extensive efforts to investigate the global spread of AIVs based on intercontinental bird flyways [8,9,10,11,12,13], one of the primary challenges is to study the role played by migratory birds in short-distance regional/provincial movements. This represents a fundamental knowledge gap in the field of avian influenza research.
Birds that reside in wetlands and aquatic environments, such as Anseriformes and Charadriiformes, are known to be natural reservoirs of AIVs [14,15]. Given that many bird species coexist in these habitats during their migration, the transmission of AIVs through the faecal-oral route via surface waters is highly probable [16]. To proactively detect the introduction of AIVs into new areas, active surveillance of bird infections is necessary to provide an early warning. To combat the growing threat of avian influenza, numerous studies have been conducted to determine the prevalence of various AIVs in different bird species and localities. For instance, Olsen et al. reviewed the worldwide occurrence of influenza A viruses in wild birds, including ducks, gulls, terns, and waders [17]. Pawar et al. estimated the prevalence of H5N1 in wild birds in India [18]. Bi et al. collected and isolated influenza H5N1 viruses from ill or deceased birds in the Sanmenxia Reservoir Area of China in 2015 [19]. Despite these efforts, there is still insufficient systematic research on influenza A (H7N9) virus across all bird species throughout the country.
In 2005, the United States Congress was urged to establish a global network for AIVs among wild birds to encourage worldwide surveillance. Since then, various genetic sequence databases, including the Global Initiative on Sharing All Influenza Data (GISAID, https://www.gisaid.org/), have been developed to facilitate the sharing of all influenza virus type sequences. Through phylogenetic analysis, genomic characterization and molecular evolution of the circulating AIVs can be uncovered [20, 21, 22, 23] using coalescent theory [24, 25]. By assuming different parametric models on population dynamics [26, 27, 28, 29, 30], the demographic history of a host population, such as the effective population size, can be reconstructed from a phylogenetic tree. Additionally, high-throughput sequencing technology makes it possible to integrate genetic variability and evolution of AIVs with virus-host ecology, such as migratory birds. In this regard, many phylogeographic studies have been conducted to examine the relationship between the global spread of AIVs and migratory bird flyways [10, 31, 32, 33]. However, because most viral sequences isolated in GISAID are not precisely annotated with geographic location and specific virus-host information, such studies can only be performed at a coarse-grained scale.
In a recent paper, researchers set out to identify the bird species and geographic hotspots relevant to the spread of influenza A (H7N9) at a finer-grained scale in China. The study collected all gene sequences of H7N9 isolated in chickens in China from 2013 to 2017, and constructed phylogenetic trees using MEGA software to explore their evolutionary relationship in terms of geographical locations.
To investigate the relationship between the identified H7N9 cases and the spatiotemporal distribution of migratory bird species, the researchers adopted the cross correlation function (CCF) based on 157,272 observation data of 1,145 bird species in China. They identified a list of possible bird species relevant to the isolated influenza A (H7N9) cases in six municipality/provinces with high incidences and explored and visualized geographical hotspots of those identified bird species.
The study’s findings have important implications for active surveillance in bird flu intervention and control. Identifying the bird species and geographical hotspots of H7N9 can provide evidence for early warning and active surveillance of bird flu in those regions. The study demonstrates the potential of using genetic sequence data and spatiotemporal distribution data of bird species to investigate the spread of avian influenza and highlights the need for systematic research on H7N9 in all bird species across the country.
Methods
Data collection and pre-processing
The Global Initiative on Sharing All Influenza Data (GISAID) provided us with full or partial hemagglutinin (HA) and neuraminidase (NA) sequences of influenza A (H7N9) virus isolated from chickens in China between January 1, 2013, and December 31, 2017. Each sequence was associated with a unique ID, annotated with a location (municipality or province) and the date of isolation. We selected HA and NA subtypes with lengths equal to or greater than 1683 and 1398, respectively, to construct our phylogenetic trees. After removing any duplicated sequences, we were left with 495 sequences for further analysis. The 495 aligned sequences are included in Additional file 2.
The citizen science project collected the bird observation data from a large number of bird enthusiasts and experts who shared their observations through an online forum. In total, 1145 bird species were observed in China, and each record included the scientific name of the bird, its locality (longitude and latitude), date of observation, number of observed birds, and the observer’s name. All observation records were reviewed by bird experts based on the characteristics of each bird species and were published in the China Bird Report annually. To avoid duplication, only one record with the highest number of observed birds was retained for each location and date. After removing the duplicates, the dataset consisted of 157,272 observation records from the years 2008 and 2009. A total of 150 migratory bird species were selected from the 1145 bird species for cross-correlation function (CCF) analysis with the assistance of bird experts (see Additional file 3). The observation data were spatially aggregated by municipality/province and temporally aggregated by weeks in a year.
Phylogenetic analysis
The hemagglutinin (HA) and neuraminidase (NA) sequences of the influenza A (H7N9) virus were first aligned using the CLUSTAL W algorithm from Clustal v.2.1 (http://www.clustal.org/download/current/) [34]. After the alignment, the HA and NA fragments were adjusted to be the same length using BioEdit v.7.0.5 (http://www.mbio.ncsu.edu/bioedit/bioedit.html) [35] and merged in MEGA v.6.0 (https://www.megasoftware.net/mega.php) [36]. The phylogenetic trees for the spliced HA and NA sequences were then constructed using three approaches: neighbour-joining (NJ), maximum parsimony (MP), and maximum likelihood (ML). The nucleotide substitution model was determined by the Akaike Information Criterion (AIC) using jModelTest v.2.1.10 (https://github.com/ddarriba/jmodeltest2) [37]. The General Time Reversible model, assuming a rate variation across sites according to a gamma-shaped distribution with invariant sites, was selected based on the dataset. The NJ approach used composite maximum likelihood algorithms to estimate the transversion/transition bias and nucleotide substitution patterns, while the ML approach employed a heuristic searching strategy for the best topology, starting with five random BioNJ trees. Tree reliabilities were tested using 1000 bootstrap replicates to produce a majority consensus tree. To simplify the reconstructed phylogenetic tree, branches with bootstrap values below 0.6 were filtered out, leaving 184 gene sequences that were reconstructed using the NJ, MP, and ML approaches. A date-calibrated tree was also generated to reveal the epidemic waves of influenza A (H7N9). Finally, the NJ tree was edited, coloured, and visualized using FigTree v.1.4.3 (https://www.megasoftware.net/mega.php) and iTOL (iTOL: http://itol.embl.de/).
Cross correlation analysis
The cross correlation technique is a statistical method used to assess the similarity between two series based on their displacement. In this study, the authors employed the sample cross correlation function (CCF) to determine the lags in observed migratory bird species that could serve as useful predictors of influenza A (H7N9) outbreaks. A positive lag value indicated a correlation between the number of observed bird species at a specific time (t) and the number of influenza A (H7N9) cases at time t + lag. The CCF command in the R software package was used to perform this analysis on bird observation data collected from six municipality/provinces, including Shanghai, Jiangsu, Zhejiang, Fujian, Jiangxi, and Guangdong, which had high incidences of H7N9 infections. These areas were chosen due to their geographical proximity to each other and the fact that the majority of the isolated H7N9 cases in chicken in China came from these regions. The identification of bird species and hotspots in these areas at a finer-grained scale would aid in the implementation of active surveillance on potential influenza A (H7N9) epidemics. The hotspots in specific municipality/provinces were identified using the kernel method in ArcGIS v10.5.
Results
Figure 1 displays the phylogenetic tree that was constructed using the NJ approach, based on 184 HA and NA segments of influenza A (H7N9) virus intercepted in China between January 1, 2013 and December 31, 2017. The name abbreviation of each sequence is used as the label for each leaf, where the first two letters represent the isolated municipality/province, and the last two numbers indicate the year of isolation. The sequences that were isolated from the same municipality/provinces are coloured similarly. The branches show bootstrap values greater than 0.5. It is noticeable that most sequences that were isolated from the same municipality/province and year are clustered together (for example, the sequences that were isolated in Guangdong with brown colour in Figure 1). Please note that we have only used the NJ tree for illustration in this paper, and similar results can be obtained using the MP and ML approaches (refer to Additional files 5 and 6).
The NJ approach was used to construct a phylogenetic tree based on 184 HA and NA segments of avian influenza A (H7N9) virus. The label of each leaf consists of a name abbreviation, where the first two letters indicate the isolation location, and the last two numbers indicate the isolation year. The sequences isolated from the same municipality/provinces are highlighted in the same colour. The branches with bootstrap values greater than 0.5 are displayed in the tree.
Influenza A (H7N9) is a highly pathogenic avian virus that has caused concern in recent years due to its potential to cause human infections. In a study investigating the geographic spread of the virus, a phylogenetic tree was constructed based on 184 HA and NA segments of the virus in China between 2013 and 2017. The tree revealed three major epidemic waves in chicken in Southeast China.
The first wave occurred mainly in Zhejiang and Jiangsu in the spring of 2013. The virus then spread to Southern China and broke out in Guangdong in 2014. The third wave occurred in almost all provinces in Eastern China from 2014 to 2017, and potentially new strains of the virus emerged.
These findings are consistent with previous studies that investigated human infections of influenza A (H7N9) virus. Understanding the geographical spread of the virus is important for implementing effective surveillance and control measures. This information can also help researchers track the evolution of the virus and identify potential new strains that may pose a threat to human health.
It is important to continue monitoring the spread of influenza A (H7N9) virus, particularly in areas where outbreaks have occurred in the past. The development of effective vaccines and antiviral drugs is also crucial in preventing and treating infections. The findings of this study provide valuable insights into the geographic spread of the virus and can inform future research and public health efforts.
The geographical spread of H7N9 virus has been a concern in Southeast China, where migratory birds are believed to be playing a crucial role. To explore this relationship, researchers conducted a cross-correlation function (CCF) analysis on 150 common migratory bird species that could potentially be associated with the identified H7N9 cases in chicken in China. The bird species were selected with the help of bird experts, and their correlations and corresponding lags were explored.
The CCF analysis results are presented in Figure 2, which illustrates the correlation coefficients for each bird species in Shanghai, Jiangsu, Zhejiang, Fujian, Jiangxi, and Guangdong. The analysis was based on time series of bird observation records with different positive lag values, where the lags were measured by weeks. The values of correlation coefficients greater than or equal to 0.27 were shown in different colors, while corresponding bird species with positive lags (i.e., Lag ≤ 10) were displayed on the x-axis.
By conducting this analysis, the researchers identified a list of migratory bird species for each municipality/province, which may be responsible for the introduction of influenza A (H7N9) epidemics in these areas. The results of the CCF analysis and the temporal distribution of identified bird species in Jiangxi Province were found to be different from those in other municipalities/provinces. The reason for this difference is that, in addition to bird migration, poultry trading is also one of the most important reasons for the geographical spread of H7N9 virus.
Overall, the findings of this study provide valuable insights into the potential mechanisms of H7N9 virus spread in Southeast China. The identification of migratory bird species associated with the spread of the virus could lead to targeted surveillance and control measures in areas at risk of outbreaks. Furthermore, the results suggest that more comprehensive investigations are needed to explain the observed differences in the temporal distribution of bird species in different municipalities/provinces.
Influenza A (H7N9) virus has been a significant public health concern since its emergence in 2013. As migratory birds are thought to be a key factor in the spread of the virus, a study was conducted to explore the correlations between the presence of the virus in chickens and migratory bird populations in six provinces in Southeast China: Shanghai, Jiangsu, Zhejiang, Fujian, Jiangxi, and Guangdong.
The study involved the analysis of 150 common migratory bird species to determine their correlations and corresponding lags with the occurrence of H7N9 cases in chicken in China. The cross-correlation function (CCF) analysis was used to identify bird species that may be responsible for the introduction of H7N9 epidemics in these areas. The analysis was conducted on time series of bird observation records with different positive lag values, measured in weeks.
The results of the CCF analysis are illustrated in Figure 2, which shows the values of correlation coefficients greater than or equal to 0.27 in different colors. The x-axis demonstrates the bird species with positive lags (Lag ≤ 10) for each municipality/province. Based on the results, a list of migratory bird species was identified for each province that may be responsible for the introduction of H7N9 epidemics in these areas.
It can be observed from the figures that the CCF results and temporal distribution of identified bird species in Jiangxi Province are different from other municipalities/provinces. This is because poultry trading is also one of the most important reasons for the geographical spread of influenza A (H7N9) virus in the region. Therefore, further investigation is required to explain the differences in the results.
Overall, the study suggests that migratory birds play a significant role in the spread of H7N9 virus in Southeast China, and identifying the bird species responsible for introducing the virus in each province can help in developing targeted prevention and control measures.
The data of influenza A (H7N9) cases and observations of migratory birds with positive lags (Lag ≤ 5) in Shanghai and other five provinces are presented in a time series format. The data of both H7N9 cases and migratory bird observations are combined and mapped over one year based on their isolation and observation dates, respectively. The x-axis of the graph starts from week 40.
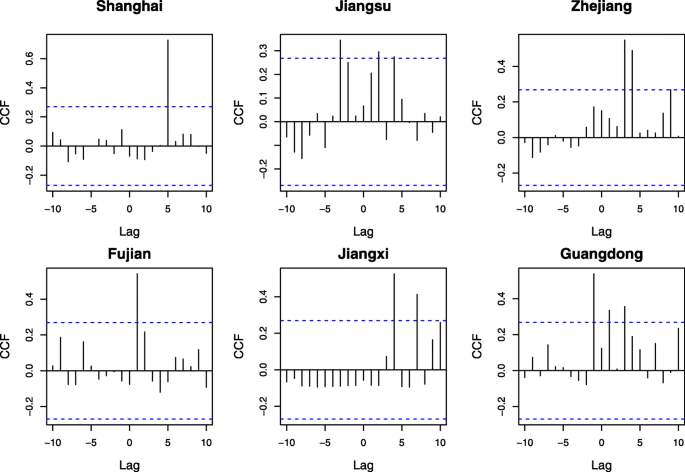
The identified bird species with positive lags (Lag ≤ 5) observation records are aggregated by weeks based on their observation dates in Shanghai and five other provinces. Figure 3 displays both time series of H7N9 cases and observation records of identified bird species, which are arranged in one year beginning from week 40. The figure suggests a significant correlation between the number of identified bird species within five weeks and the incidence of influenza A (H7N9) cases. Moreover, the CCF analysis is carried out to examine the cross-correlation between H7N9 cases and identified bird species in each municipality/province. Figure 4 illustrates the analysis outcomes, with the dotted blue line representing the correlation coefficient threshold value of 0.27. The results reveal that in each municipality/province, there is a minimum of one lag within five weeks with a correlation coefficient higher than the threshold.
In order to investigate the potential correlation between the observed migratory bird species and the spread of Influenza A (H7N9) virus in Shanghai and other five provinces, a cross-correlation function (CCF) analysis was conducted. The CCF analysis is a statistical method used to evaluate the correlation between two time series data sets, in this case, the time series of H7N9 cases and the observation records of identified bird species.
The analysis focused on identifying positive lags (Lag ≤ 5) between the time series of observed bird species and H7N9 cases. A positive Lag value indicates the correlation between the amount of observed bird species at a given time and that of H7N9 cases at a later time (t + Lag), where Lag is measured by weeks.
Figure 4 shows the results of the CCF analysis, where the dotted blue line represents the threshold value of the correlation coefficient, which is set to 0.27. The figure illustrates that in each of the six municipalities/provinces, there is at least one Lag value within five weeks with a correlation coefficient greater than the threshold. This suggests that there is a strong correlation between the number of observed bird species and the incidence of H7N9 cases within a given timeframe.
These findings are crucial in understanding the potential role of migratory birds in the spread of the H7N9 virus. The results of the analysis provide insights into identifying specific bird species that may be responsible for introducing the virus to a particular area, which can aid in implementing targeted control measures.
The distribution of identified migratory bird species and their correlation with the spread of Influenza A (H7N9) virus in Eastern China is the focus of this study. The results of the study are presented in Figs. 5 and 6. Fig. 5 illustrates the geographic distribution and kernel density of identified bird species with positive lags (Lag ≤ 5). The blue nodes indicate the number of birds observed, while the coloured surface indicates the density magnitude of bird species after smoothing. The study further investigates the potential hotspots of H7N9 epidemics based on the geographical distribution of identified migratory bird species in six municipalities/provinces in Eastern China, as shown in Fig. 6.
The study shows that most of the hotspots of identified migratory bird species are located along the coastal areas or around large lakes. For instance, hotspots can be observed in Shanghai, Jiangsu, Zhejiang, Fujian, and Guangdong along the east coast of China. In Jiangsu Province, another hotspot is located around Taihu Lake, which is close to Wuxi City and Suzhou City. In Jiangxi Province, the major hotspot is located around Poyang Lake.
The findings of this study have important implications for the prevention and control of H7N9 epidemics. By identifying the hotspots of migratory bird species, authorities can take measures to monitor and control the spread of the virus in these areas. The study highlights the importance of continued surveillance and monitoring of migratory bird species in areas prone to the introduction of H7N9 virus epidemics.
The distribution of bird species identified with positive lags Lag ≤ 5 in China is depicted on the map. The blue nodes’ size represents the total number of bird species observed, while the coloured surface indicates the density magnitude of bird species after smoothing. The map was created using ArcGIS v.10.5.
Geographical mapping is an essential tool for identifying and analyzing the spread of infectious diseases. In this regard, researchers have used ArcGIS v.10.5 to generate maps that illustrate the geographical hotspots of identified bird species with positive lags Lag ≤ 5 in Shanghai and other five provinces. The size of the nodes in blue represents the total number of identified bird species in each municipality/province, while the coloured surface represents the density magnitude of bird species after smoothing.
These maps help identify areas where the influenza A (H7N9) virus may be introduced by migratory birds. The identification of hotspots is particularly important for the early detection and control of the virus, as it can lead to the implementation of targeted prevention and control measures.
In Shanghai, Jiangsu, Zhejiang, Fujian, Jiangxi, and Guangdong, the maps show that most hotspots are located along the coastal areas or around large lakes. For example, there are hotspots in Shanghai, Jiangsu, Zhejiang, Fujian, and Guangdong that are along the east coast of China. In Jiangsu Province, there is another hotspot that is around Taihu Lake and close to Wuxi City and Suzhou City. In Jiangxi Province, the major hotspot is located around Poyang Lake.
Overall, the geographical hotspots of identified bird species with positive lags Lag ≤ 5 provide critical insights into the spread of the H7N9 virus in Eastern China along the East Asian-Australasian flyway. These maps help in the development of effective strategies for controlling the spread of the virus and reducing the risk of outbreaks in both humans and animals.
Discussion
In order to effectively monitor and identify avian influenza viruses (AIVs) in China, it is crucial to systematically identify relevant bird species and their geographical hotspots at a more detailed level. One of the key challenges in achieving this is understanding the migratory patterns of various bird species. Traditionally, bird banding has been a widely-used method to track bird movements and life history, but it is expensive and time-consuming. Additionally, only a small percentage of bird rings can be recycled. GPS technologies have also been utilized for satellite tracking, but it is costly and cannot be widely applied to monitor a large number of bird species.
In recent years, citizen science has emerged as a promising method for collecting vast amounts of bird observation data across large areas through crowd-sourcing data collection platforms. For example, the Cornell Lab of Ornithology created eBird and several mobile applications to collect and share bird observation data from around the world. Similarly, in China, bird-watching enthusiasts have established the China Bird Watching Network and share their observations through the Bird Record Center of China. The China Ornithological Society has also published annual reports on bird records since 2003, known as the China Bird Report.
Through these crowd-sourced data collection platforms, it is possible to analyze the spatiotemporal distribution and migration patterns of bird species at a finer-grained scale and within a larger geographical area. This approach provides a more cost-effective and efficient way to systematically identify relevant bird species and their geographical hotspots, which is crucial for active surveillance of AIVs in China.
With the advancement of high-throughput sequencing technology, it has become easier to isolate gene sequences of avian influenza viruses (AIVs). The Global Initiative on Sharing All Influenza Data (GISAID) provides a platform for sharing AIV gene sequences with additional information, such as provincial location and isolation date. Several phylogeographic studies have investigated the global spread of AIVs, such as H5N1. However, due to the lack of precise location coordinates and species-specific information associated with avian isolates in GISAID, phylogeographic analysis can only be conducted at a coarse-grained scale. To enable more comprehensive phylogeographic studies, it is important to provide accurate information about avian influenza isolates. In this study, key migratory birds from 1145 bird species in China were identified, and their spatiotemporal distribution can serve as a prior for date calibration and location estimation in Bayesian phylogeographic methods. This information can help to understand the spread of AIVs at a finer-grained scale.
During breeding or winter seasons, bird species tend to gather at stopovers or habitats with high population density, which can lead to transmission of AIVs among different migratory bird species and between migratory and domestic fowls, causing new infections in different locations. These gathering locations are likely to be the ancestral sites where reassortment or recombination of different AIVs occur. To predict and monitor the spread, recombination, and reemergence of AIVs among migratory birds, the crowd-sourcing data collection platform for bird species all over China provides a new perspective. With 157,272 observation records of 1,145 bird species collected in China, an in-depth investigation was conducted on the relationship between the spatiotemporal distribution of migratory birds and historical epidemics of influenza A (H7N9). Key bird species and their geographical hotspots were identified for the implementation of active surveillance of influenza A (H7N9) in high epidemic areas. In the future, it is expected to integrate bird observation data in China with more comprehensive records from eBird to conduct a thorough investigation about AIVs in birds all over the world. With the joint efforts of ornithologists, epidemiologists, and molecular biologists, the potential intercontinental movement of influenza A (H7N9) virus can be assessed, as well as the possible introduction pathway of novel AIVs across continents.
This paper has some limitations due to the current availability of data. Firstly, only the relationship between migratory birds’ spatiotemporal distribution and the spread of influenza A (H7N9) cases in chicken in China was explored. It would be more informative to investigate the relationship between bird migration and the spread of other AIVs (e.g., H5N1). Secondly, the observation data on birds is imbalanced in China, with more records along the Eastern China flyway than the Central Asian flyway. However, this does not necessarily mean that there are more bird species in the former region. More data can be collected in the future with the help of citizen science, mitigating the impact of data imbalance. Thirdly, geographical hotspots were only identified in six municipalities/provinces, representing 91.2% of isolated H7N9 cases in chicken in GISAID. A more systematic analysis on different types of AIVs in China is needed by integrating molecular evolution of AIVs and spatial ecology of migratory birds. Gene sequences of avian isolates should be annotated with specific bird names and GPS locations when uploaded to the GISAID database. Finally, while key bird species and geographical hotspots were identified based on CCF analysis, it does not necessarily mean that H7N9 cases in those locations were introduced by migratory birds. Poultry trading data should be considered in future analyses.
Conclusions
In this study, we have examined the correlation between the spread of influenza A (H7N9) outbreaks and the temporal and spatial distribution of bird species in China. Our analysis has identified key migratory bird species and geographical hotspots associated with the introduction of H7N9 epidemics in six major areas in China. To achieve this, we conducted phylogenetic analysis on the HA and NA segments of the virus isolated in chicken in China from 2013 to 2017. This allowed us to identify three major epidemic waves of H7N9 in Eastern China along the East Asian-Australasian flyway. Furthermore, we collected more than 157,000 bird observation records from a crowd-sourced data collection platform, and through cross-correlation analysis, identified 150 migratory bird species whose temporal distribution strongly correlated with H7N9 cases within five weeks. By mapping these identified species’ spatial distribution, we were able to determine potential hotspots of H7N9 epidemics in Shanghai, Jiangsu, Zhejiang, Fujian, Jiangxi, and Guangdong. Our findings could aid public health authorities in implementing active surveillance and control measures during AIV epidemics.
Abbreviations
The Akaike Information Criterion (AIC)
Avian influenza virus, also known as bird flu, is a highly contagious viral disease that affects birds and can also spread to humans. It is caused by influenza A viruses that primarily affect birds, but can also infect humans and other mammals. In birds, the virus can cause mild to severe illness, and in some cases, death.
The avian influenza virus can be transmitted through contact with infected birds or their secretions, contaminated equipment or surfaces, and contaminated food or water. It can also be spread through the air by infected birds, especially during migratory flights.
The symptoms of avian influenza in birds include respiratory distress, decreased egg production, and sudden death. In humans, the symptoms are similar to those of other types of influenza, including fever, cough, sore throat, muscle aches, and fatigue. However, in severe cases, the virus can lead to respiratory failure, multi-organ failure, and death.
The avian influenza virus has the potential to cause pandemics in humans, as was seen in 2009 with the H1N1 pandemic. Therefore, it is essential to monitor the virus and develop effective prevention and control strategies.
To prevent the spread of avian influenza virus, it is crucial to practice good hygiene, avoid contact with infected birds and their secretions, and cook poultry thoroughly. Vaccines are also available for some strains of the virus, but they are not effective against all strains.
In conclusion, avian influenza virus is a significant public health concern that can have devastating consequences for both birds and humans. It is crucial to remain vigilant and take appropriate measures to prevent its spread and minimize its impact.
The function of cross-correlation is to measure the similarity between two sets of data, where one set is shifted relative to the other. It can be used to determine if there is a relationship between two variables and to quantify the strength and direction of that relationship.
The function of cross-correlation is to measure the similarity between two sets of data, where one set is shifted relative to the other. It can be used to determine if there is a relationship between two variables and to quantify the strength and direction of that relationship.



